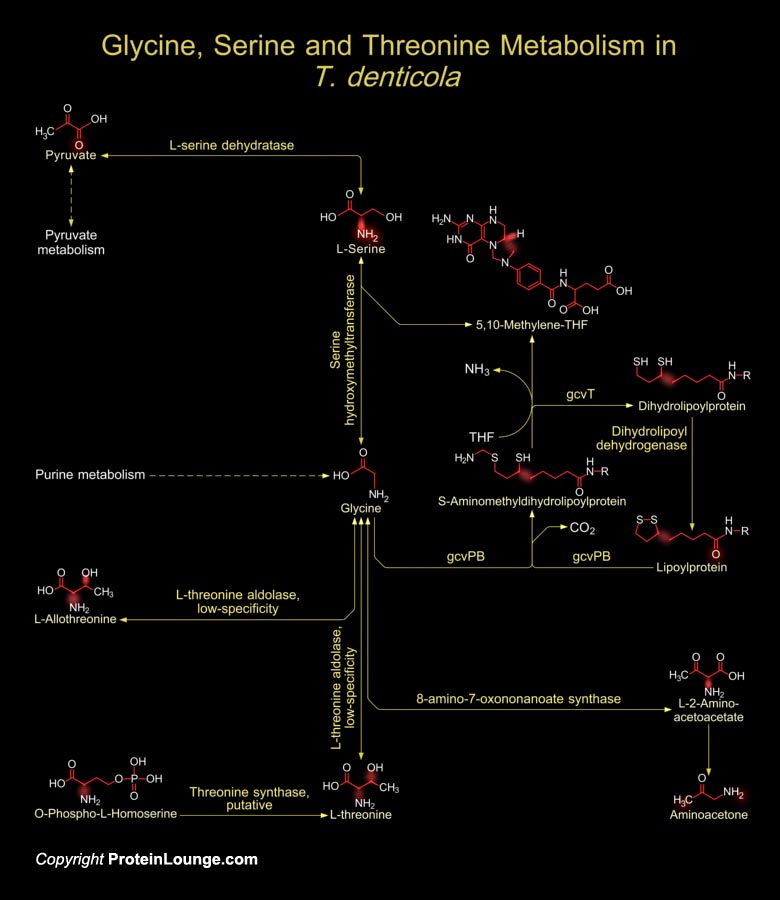
The Gram-negative oral spirochete T. denticola (Treponema denticola) is predominantly associated with the incidence and severity of human periodontal disease. T. denticola is an obligate anaerobe commonly associated with an inflammation of gum tissue that frequently precedes bone resorbtion and subsequent tooth loss of humans. T. denticola lacks a traditional LPS (Lipopolysaccharide), as do many other Spirochetes, for which these are resistant to cationic peptide antibiotics as well as to human Beta-Defensins, as these interact strongly with LPS due to its negative charge. Such a mechanism prompts T. denticola to thrive in gum tissue and affect substrate specificity and permit the utilization of amino acids and other compounds for substrate phosphorylation and ATP[..]
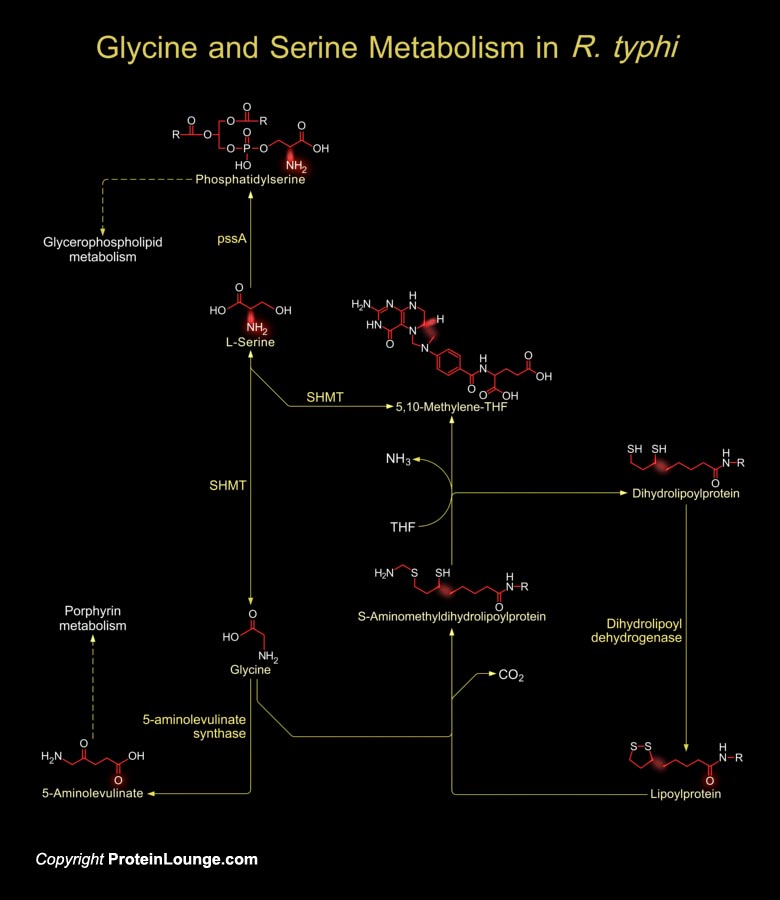
Rickettsial infections have played a significant role in the history of Western civilization through epidemic Typhus and Rickettsia used arthropod vector for the spread of Typhus fever. R. typhi (Rickettsia typhi) is a smaller in size than normal bacteria and obligately intracellular pathogen. However, they are true bacteria, small coccobacilli that are normally stained with Giemsa and poorly by the Gram stain. Its cell wall morphology is that of a Gram-negative bacillus. Phylogenetically a member of the Alpha-subgroup of Proteobacteria, R. typhi along with R. prowazekii (Rickettsia prowazekii) is considered to be Typhus group Rickettsia (Ref.1). R. typhi evolved in close association with its arthropod vectors, various species of fleas. In its best known zoonotic[..]
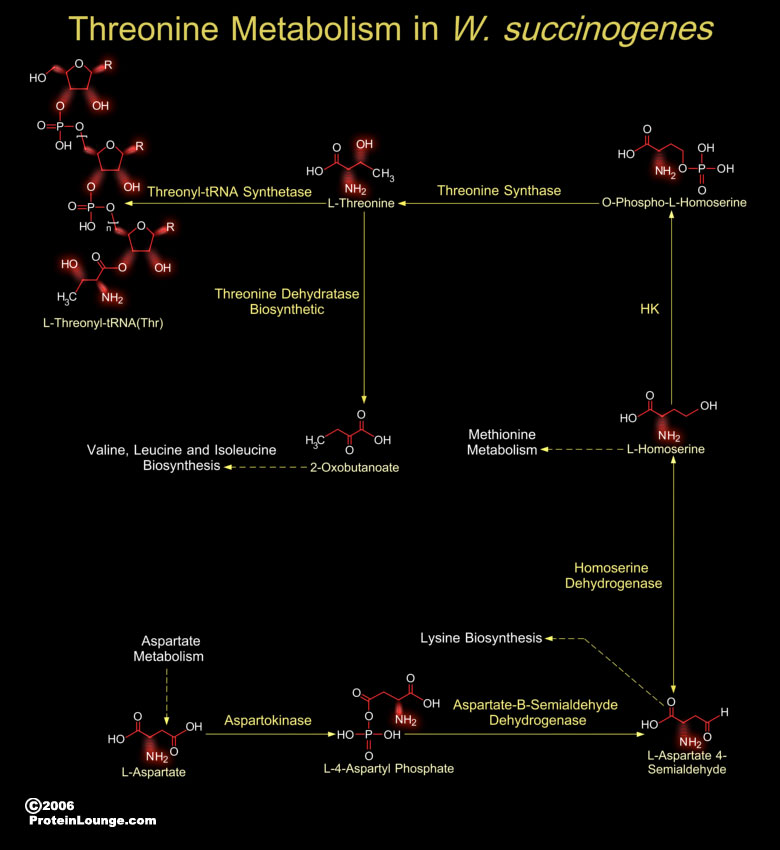
W. succinogenes (Wolinella succinogenes) is a non-fermenting bacterium that grows anaerobically. It is rod-shaped bacteria with monotrichous flagellation and the insertion of the flagellar motor is into the pole of the cell. W. succinogenes resides in the bovine rumen, the human gingival sulcus, and dental root canal infections. W. succinogenes, is closely related to the pathogenic bacteria H. pylori (Helicobacter pylori) and C. jejuni (Campylobacter jejuni) (Ref.1). Despite being considered non-pathogenic to its bovine host, W. succinogenes holds an extensive repertoire of genes homologous to known bacterial virulence factors. Many of these genes have been acquired by lateral gene transfer. W. succinogenes genome also reveals genes related to soil and plant associated[..]
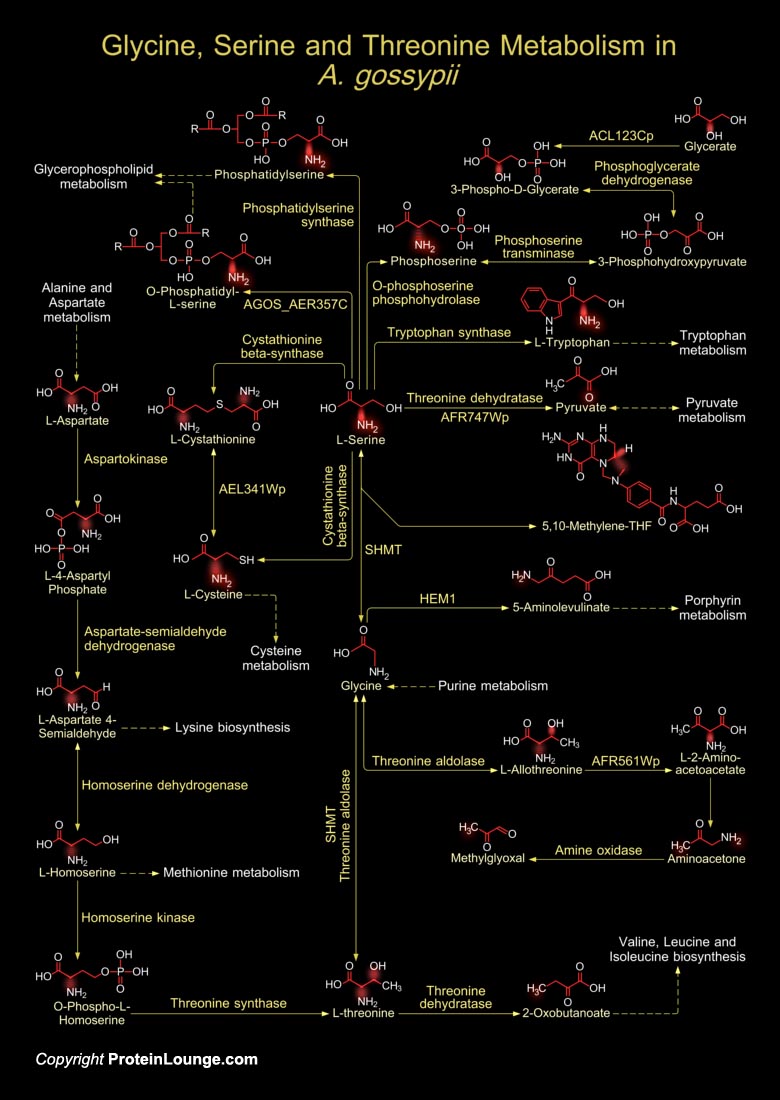
The yeast A. gossypii (Ashbya gossypii), also known as E. gossypii (Eremothecium gossypii), is a free-living hemiascomycete that utililizes amino acid metabolisms to obtain its nitrogen source. For such reason the metabolism of Glycine, Serine and Threonine is essential for the survival of A. gossypii. Yeast physiology can be either obligately aerobic or facultatively fermentative. There is no known obligately anaerobic yeast. Yeasts generally use L-Serine or L-Threonine as sole nitrogen source. (Ref.1 & 2). L-Threonine is generally synthesized from Pyruvate and L-Aspartate. The enzyme Threonine Dehydratase converts Pyruvate (from Pyruvate metabolism) to amino acids like L-Serine. SHMT (Serine Hydroxymethyltransferase) then catalyzes the cleavage of L-Serine to[..]
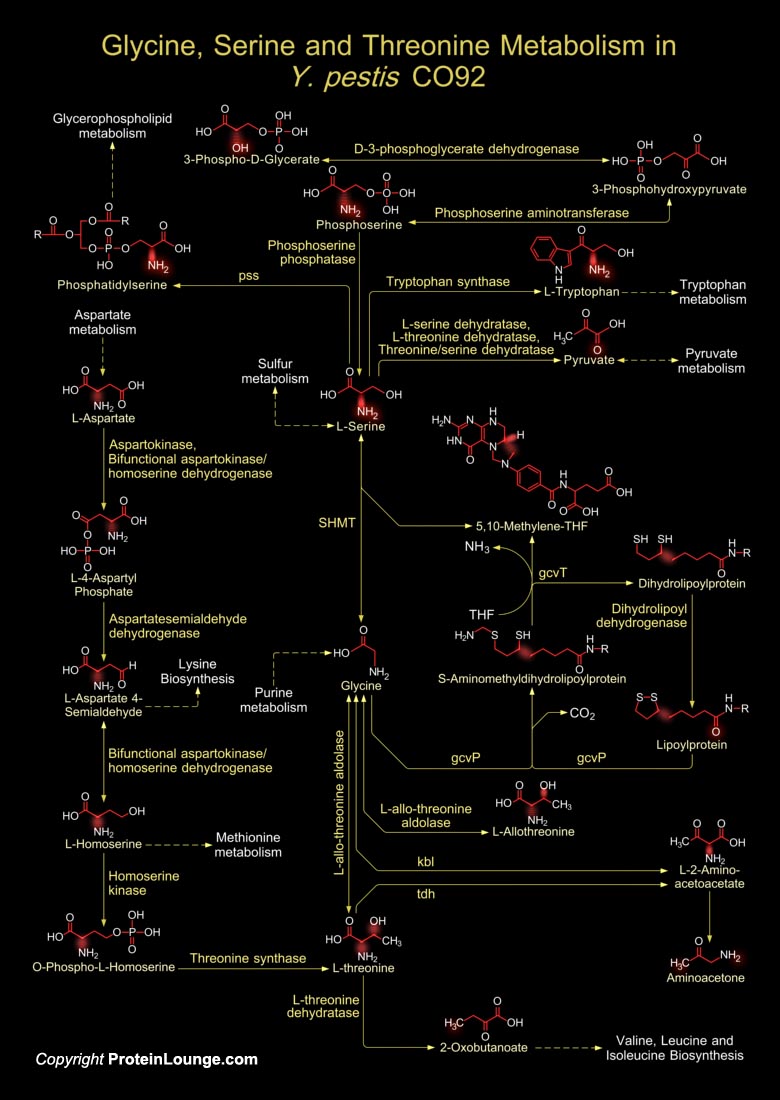
Y. pestis (Yersinia pestis), the Gram-negative Coccobacillus belonging to the Enterobacteriaceae is the causative agent of Plague and is arguably the deadliest pathogen in history. It is one of several agents likely to be used as a biological weapon in a bioterrorism event. Yersinia sp. is responsible for disease syndromes ranging from Gastroenteritis to Plague. Y. pestis is actually categorized into three subtypes or biovars; Antiqua, Medievalis, and Orientalis, each associated with a major pandemic. Y. pestis strand CO92 is in the biovar Orientalis. The biovar Orientalis bacteria are responsible for the modern Plague. Y. pestis CO92 has a circular chromosome that is 4.65Mb in size. Strain CO92 has one less rRNA operon than strain KIM (Ref.1). Y. pestis is rod shaped[..]
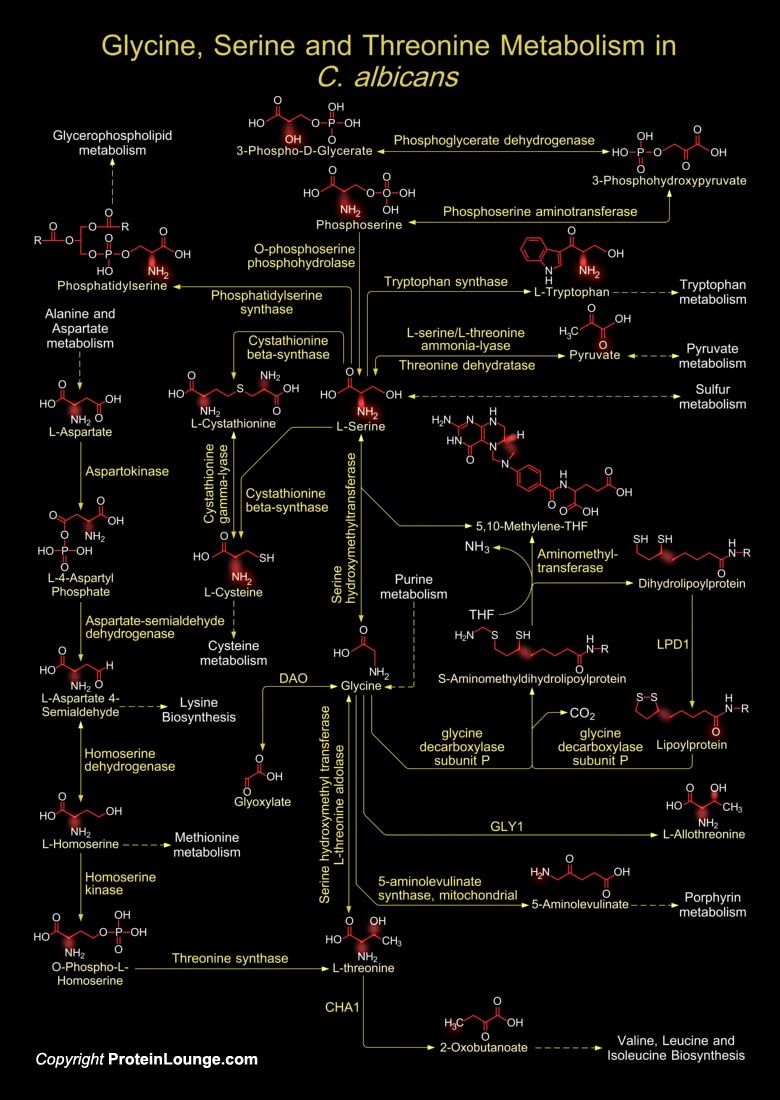
Like many bacteria, yeast species can form biofilms on several surfaces. C. albicans (Candida albicans) colonizes in the surfaces of catheters, prostheses, and epithelia, forming biofilms that are extremely resistant to anti-fungal drugs. The protein Gcn4 (Transcriptional Activator Gcn4), a regulator of amino acid metabolism, is required for normal biofilm growth. The biochemical mechanisms including activation of the sulfur-amino acid biosynthesis pathway is a feature of C. albicans biofilms. For such reasons the metabolism of Glycine, Serine and Threonine is of vital importance as it regulates morphogenesis and virulence in C. albicans. Yeasts also use L-Serine or L-Threonine as sole nitrogen source for their survival. L-Threonine is generally synthesized from[..]
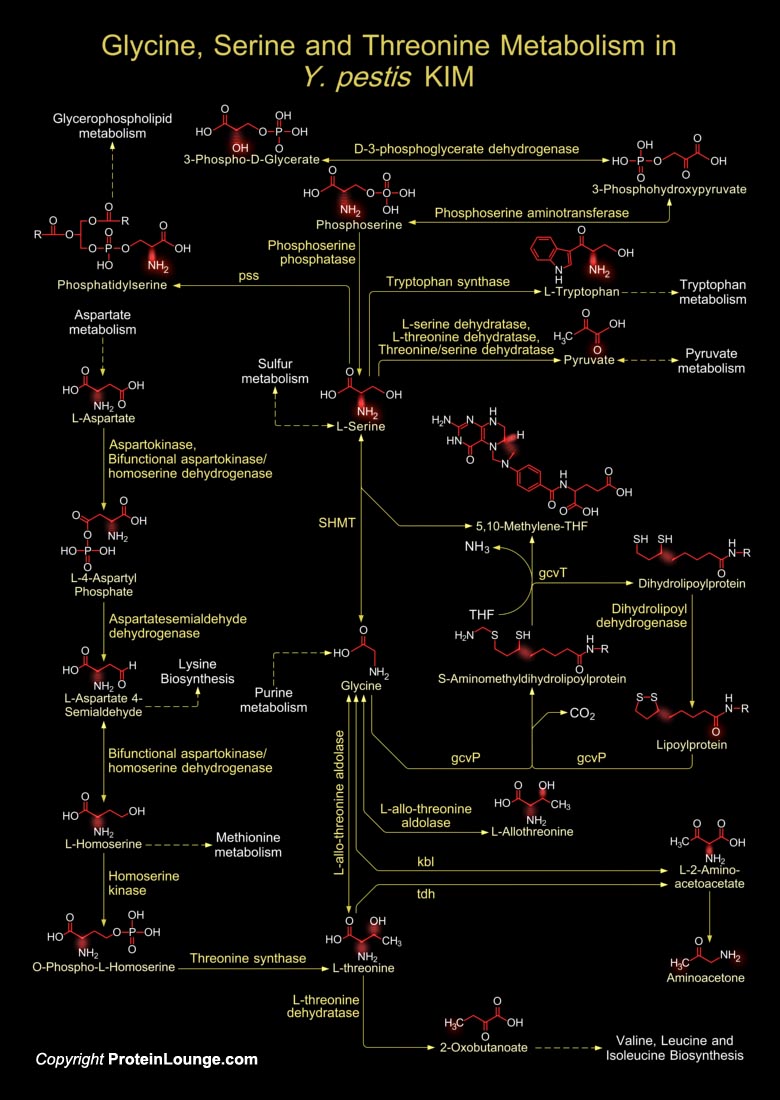
Y. pestis (Yersinia pestis) strand KIM belongs to biovar Mediaevalis. Y. pestis is actually catagorized into three subtypes or biovars; Antiqua, Medievalis, and Orientalis, each associated with a major pandemic and it is believed that Y. pestis is a clone that evolved from Y. pseudotuberculosis (Yersinia pseudotuberculosis) about 1.5 to 20 thousand years ago. Biovar Mediaevalis is thought to have descended from the bacteria that caused the second pandemic, the Black Death. The Black Death came in three forms, the bubonic, pneumonic, and septicemic. Each different form of plague killed people in a vicious way. All forms were caused by a bacterium called Y. pestis. This Coccobacillus is rod shaped, Gram-negative, and non-motile but has two distinct flagellar gene[..]
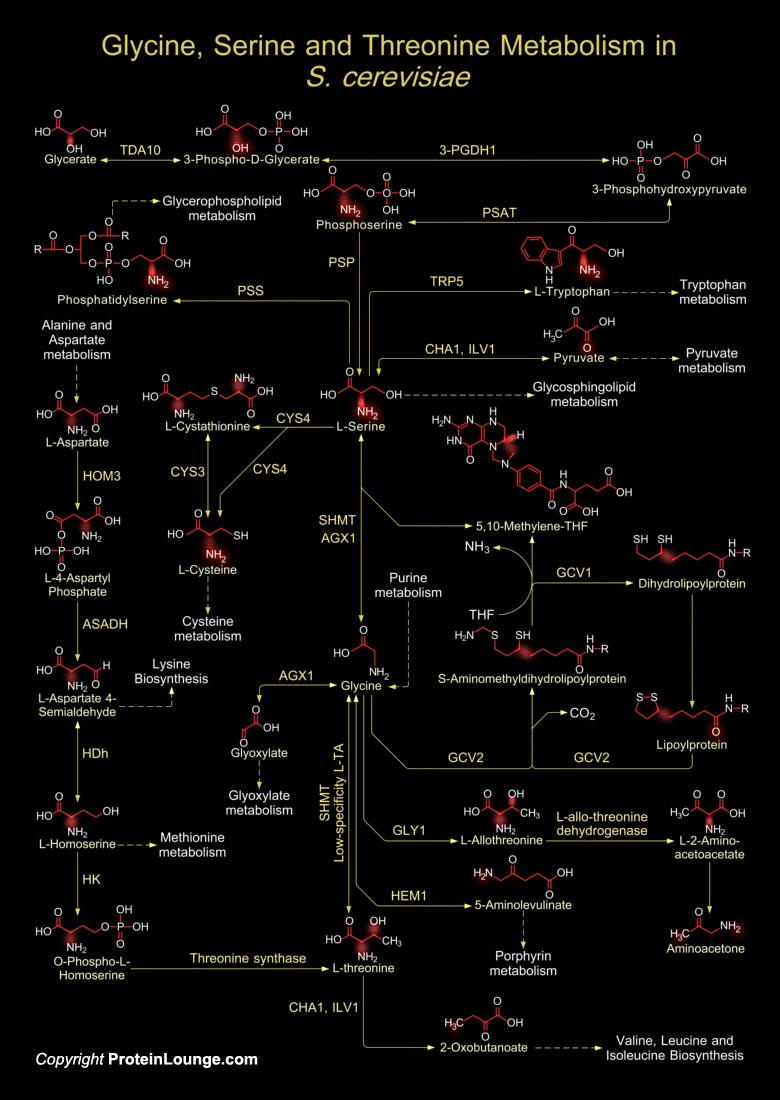
S. cerevisiae (Saccharomyces cerevisiae) are single-celled fungi which multiply by budding or in some cases by division (fission). Yeast fermentations comprise the oldest and largest application of microbial technology. Yeast physiology can be either obligately aerobic or facultatively fermentative. There is no known obligately anaerobic yeast. In pharmacy and chemistry, a large number of substances (Vitamins and enzymes) are extracted from this organism. In addition, some medications are now produced by manipulated yeasts, for example vaccines against Hepatitis-B surface antigen are produced by S. cerevisiae utilizing recombinant technology. The metabolism of Glycine, Serine and Threonine in S. cerevisiae is of vital importance as they use L-Serine or L-Threonine as[..]
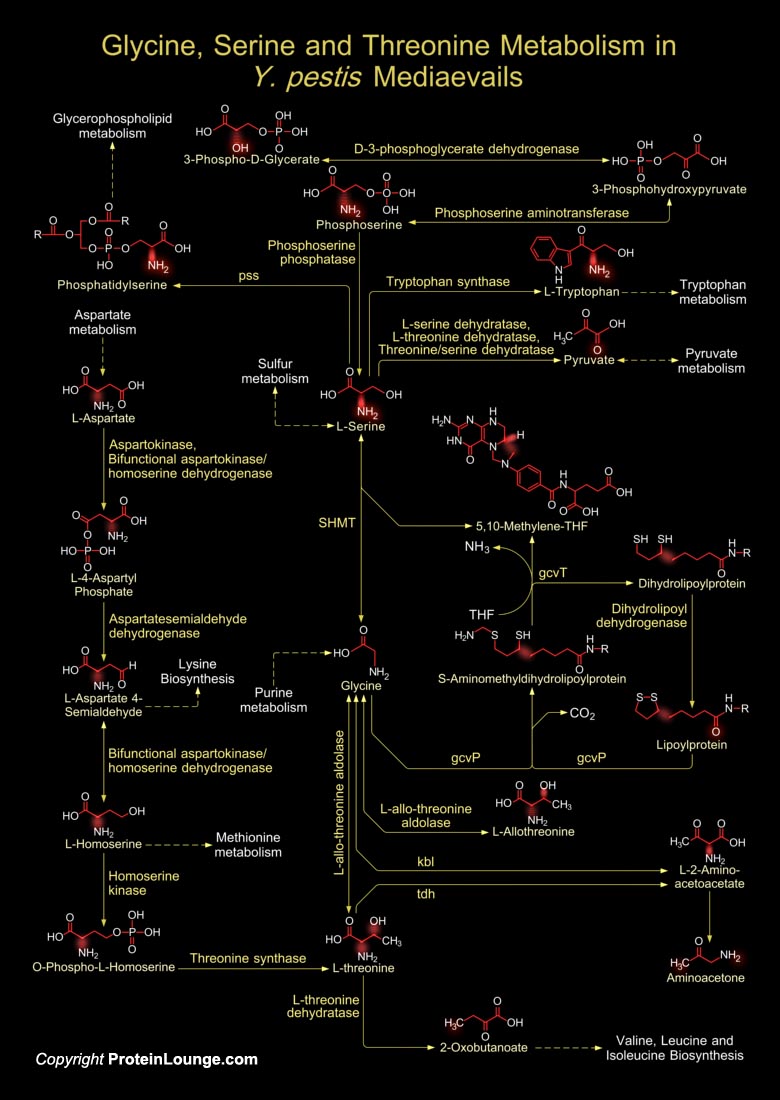
Y. pestis (Yersinia pestis) is rod shaped, Gram-negative, and non-motile but has two distinct flagellar gene clusters; one set is incomplete and the other produces a truncated protein, which acts as a transcriptional activator for the flagellar genes. Y. pestis, a Group-A bioterrorism agent, causes Plague, a re-emerging zoonotic disease transmitted to humans through flea bites and typically characterized by the appearance of a tender and swollen lymph node, the bubo. It is believed that Y. pestis is a clone that evolved from Y. pseudotuberculosis (Yersinia pseudotuberculosis) about 1.5 to 20 thousand years ago. Y. pestis is primarily a rodent pathogen, with humans being an accidental host when bitten by an infected rat flea (Ref.1). The rodents’ fleas, such as[..]
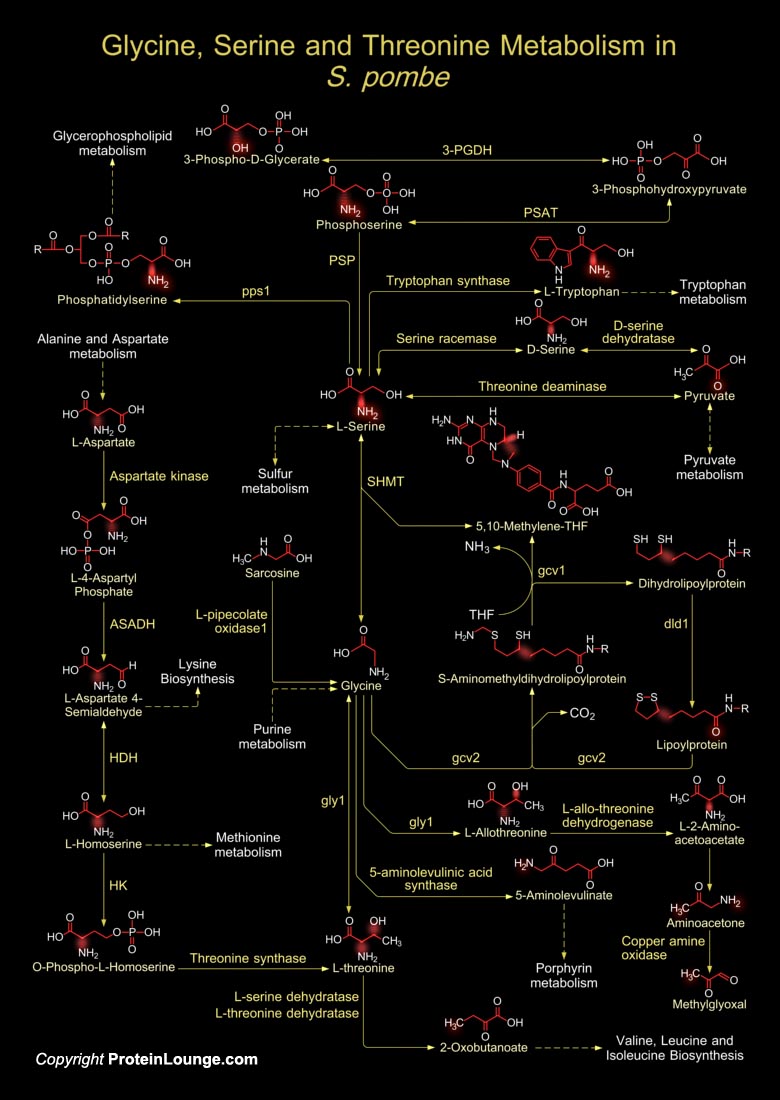
The metabolism of Glycine, Serine and Threonine is essential for the unicellular eukaryote S. pombe (Schizosaccharomyces pombe), as it uses L-Serine or L-Threonine as sole nitrogen source for its survival. During the metabolism of these amino acids, L-Threonine is generally synthesized from Pyruvate and L-Aspartate. The enzyme Threonine Ammonia Lyase/Threonine Dehydratase converts Pyruvate (from Pyruvate metabolism) to amino acids like L-Serine. Serine Hydroxymethyltransferase/Glycine Hydroxymethyltransferase then catalyzes the THF (Tetrahydrofolate) dependent cleavage of L-Serine to Glycine and 5,10-Methylene-THF (5,10-Methylenetetrahydrofolate). Serine Hydroxymethyltransferase is the major provider of one-carbon units in the cell, which are derived from catabolism of[..]
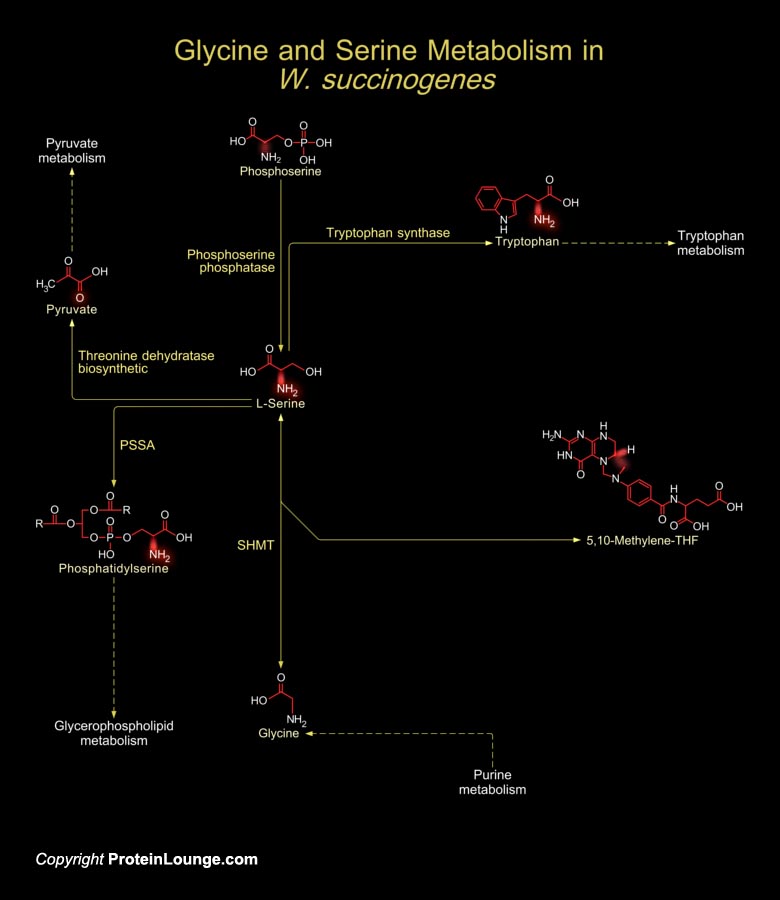
W. succinogenes (Wolinella succinogenes) is a non-fermenting bacterium that grows anaerobically. It is rod-shaped bacteria with monotrichous flagellation and the insertion of the flagellar motor is into the pole of the cell. W. succinogenes resides in the bovine rumen, the human gingival sulcus, and dental root canal infections. W. succinogenes, is closely related to the pathogenic bacteria H. pylori (Helicobacter pylori) and C. jejuni (Campylobacter jejuni) (Ref.1). Despite being considered non-pathogenic to its bovine host, W. succinogenes holds an extensive repertoire of genes homologous to known bacterial virulence factors. Many of these genes have been acquired by lateral gene transfer. W. succinogenes genome also reveals genes related to soil and plant associated[..]
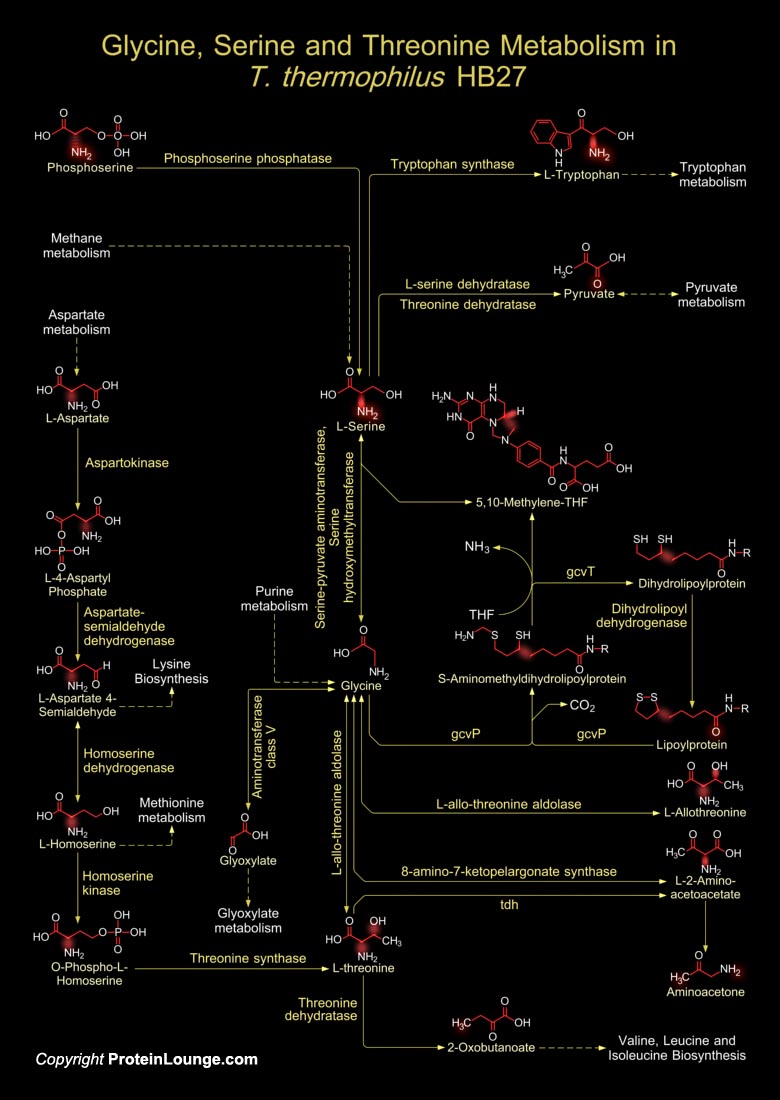
T. thermophilus (Thermus thermophilus) is a Gram-negative, aerobic eubacterium which grows at temperatures ranging from 50 to 82 degrees Centigrade. The organism, T. thermophilus strain HB27 grows in a natural thermal environment with an optimal growth at 68 degrees Centigrade and pH 7.0. Most extreme thermophiles that live in geothermal environments are strict anaerobes as a consequence of the adaptation to the low solubility of oxygen at these temperatures. However, Thermus species are an exception because they are strictly aerobic chemorganotrophs. T. thermophilus HB27 is unable to grow under anaerobic conditions but the ability to grow anaerobically by nitrate reduction can be transferred to the aerobic strain HB27 by conjugation. Thermophilic organisms, like the[..]

